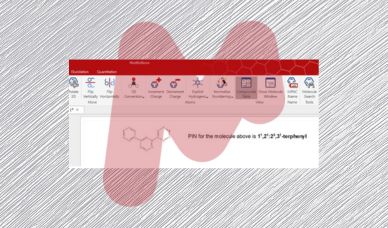
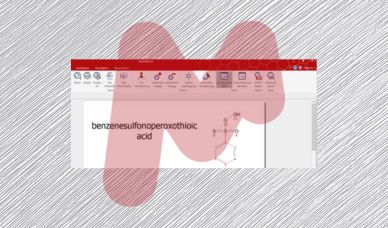
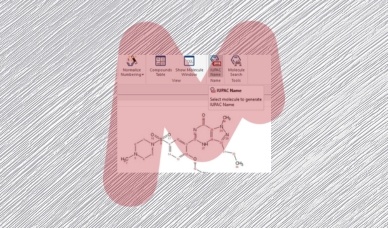
Mnova IUPAC Name algorithm – Part I
The IUPAC naming algorithm in Mestrelab Research SL’s Mnova IUPAC Name plugin represents an innovative step forward compared to other software products with similar functionality available in the marketplace.
Traditionally, naming software solutions have relied on the building of extensive ‘dictionaries’ of large groups of atoms and their corresponding naming string, and then on the concatenation of such naming strings upon analysis of the structure to extract those fragments.
Mnova IUPAC Name is different. It works with a much smaller library of functional groups, and with extensive algorithmia for their correct combination following IUPAC conventions.
One of the resulting advantages of Mestrelab IUPAC Name is its ability to generate correct IUPAC names for most of the possible modifications of suffix groups. The relevance of this development can be illustrated by comparing the results of Mnova IUPAC Name with those returned by other 3 similar programs.
Let’s look at the simplest suffix group, carboxylic acid:
In this group, =O group may be replaced with:
=N (this is carboximidic acid)
=N-NH (carbohydrazonic acid)
=S, =Se, =Te (carbothioic acid, carboselenoic acid, carbotelluroic acid)
At the same time, -OH group may be replaced with:
-SH, -SeH, -TeH (carbothioic acid, carboselenoic acid, carbotelluroic acid)
-O-, -S-, -Se-, -Te- (carboxylate, carbothioate, etc)
-N (carboxamide)
-N-NH (carbohydrazide)
-Y-YH, where Y may be any atom from list O, S, Se, Te, so there are totally 16 variants
(carboperoxoic acids, etc).
So we have 6 * 26 = 156 possible modifications of alone carboxylic acid.
Other suffix groups (sulfonic acid, sulfinic acid, selenonic acid, seleninic acid, telluronic acid,
tellurinic acid, etc) have even more possible modifications.
And MestreLab/Name generates correct IUPAC names for all of them.
The table below contains names of several modifications of carboxylic acid generated by
MestreLab/Name, and the other 3 applications. The correct IUPAC names are bolded. Their
correctness is checked according to IUPAC Blue Book 2013 (Nomenclature of Organic
Chemistry. IUPAC Recommendations and Preferred Names 2013), pages 437-438.
The following table shows a very simple example which illustrates how even simple molecules
may be misnamed by alternative approaches to the naming problem. Mestrelab’s approach is
novel and, by containing algorithmia which is capable of combining successfully much smaller
functional groups, it can avoid many of the issues experienced by other software packages.
Overall, Mnova IUPAC Naming is an ongoing project which will keep improving in the next
releases of Mnova. There are a number of classes of molecules that cannot still be named, but
we are confident that those that are named, are properly done.
You can also read some more related articles on Mnova IUPAC Name algorithm: